The first comprehensive, large-scale analysis available on the preprint server bioRxiv* attempted to identify specific genomic signatures of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) strains within and across individual hosts – revealing recombination events, specific geographic profiles, as well as multi-strain and/or superinfections characteristic for the coronavirus disease (COVID-19) pandemic.
The current COVID-19 pandemic unveiled an urgent need for accurate evolutionary and transmission history data to inform outbreak management in real-time, devise appropriate mitigation strategies and implement public health policies. Achieving this goal requires a deeper understanding of SARS-CoV-2.
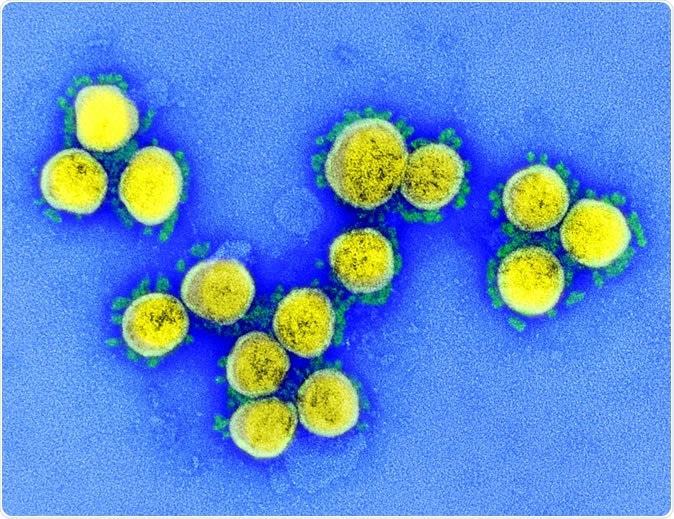
SARS-CoV-2 – Transmission electron micrograph of SARS-CoV-2 virus particles, isolated from a patient. Image captured and color-enhanced at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. Credit: NIAID
Unlike a myriad of studies that characterized viral genomes on the basis of consensus sequences, the researchers from the University of Illinois at Urbana-Champaign decided to look into the heterogeneity within individual samples and shared mutational signatures across samples.
Appraising viral diversity
This group of researchers analyzed a total of 621 SARS-CoV-2 bulk sequencing samples from the National Center for Biotechnology (NCBI) Information Sequence Read Archive to identify viral genomic signatures/strains occurring within and across COVID-19 patients.
They have also developed algorithms to deconvolve the bulk sequencing samples in the discovery set and to eventually find the proportion of the inferred strains in each Sequence Read Archive sample, that was then validated against 7,540 GISAID database of consensus sequences.
GISAID is a public-private collaboration that enables the rapid sharing of genetic sequencing data related to COVID-19 in an openly accessible database. Currently, this database contains more than 2,600 SARS-CoV-2 genomic sequences modeled in real-time, which helps significantly in detecting viral mutations and tracking its global movements.
After a detailed phylogenetic analysis of the inferred strains, the deconvolution approach enabled an in-depth look at global viral diversity below the consensus level – pinpointing in turn hosts with strains originating from distinct clades, as well as numerous examples of recombination. Mutation and immunogenicity studies helped to round the picture.
“It is critical to evaluate viral diversity below the consensus level as minor variants may impact the patterns of virulence and person-to-person transmission efficiency,” study authors explain their methodological approach.
Geographical and individual clustering
In a nutshell, this study showed evidence for within-host viral diversity across phylogenetic clades, multi-strain infections, and superinfections, putative cases of recombination events, as well as distinctive strain profiles across time and geographic locations.
More specifically, a phylogenetic analysis effort clustered the analyzed strains into four distinct clades, while spatiotemporal analysis revealed that Clade 3 arose most recently. Furthermore, the separate epidemiological analysis showed that Clade 3 has a higher average reproduction number in comparison to other clades, being most prevalent in European and North American countries.
The reproduction number represents a key concept in infectious disease epidemiology and indicates the propensity of a specific infectious agent for epidemic spread. Basically, the larger the value of a reproduction number, the harder it is to control the outbreak – which may explain why Clade 3 had such a success in Europe and the US.
These findings have paramount implications for understanding both the mutability and transmissibility of specific SARS-CoV-2 strains. In addition, the practical value of examining the viral composition of COVID-19 patients (below the consensus level) is the possibility of meticulous contact tracing.
Future research avenues
The algorithms presented in this study may be applied to analyze the viral diversity of SARS-CoV-2 at the organ level but also to correlate identified strains with outcome measures such as disease severity, morbidity, and mortality in order to get more insights into COVID-19 pathogenesis.
An exciting research avenue will be to study whether the presence of specific strains within a host comes with adverse outcomes. Moreover, the strains linked to each infected host may improve transmission history reconstruction by employing techniques that support multi-strain infections and multiple samples per host.
The end-result may be a somewhat accurate contact tracing method. “Our findings and algorithms will facilitate more detailed evolutionary analyses and contact tracing that specifically account for within-host viral diversity in the ongoing COVID-19 pandemic as well as future pandemics”, conclude study authors.
*Important Notice
bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.