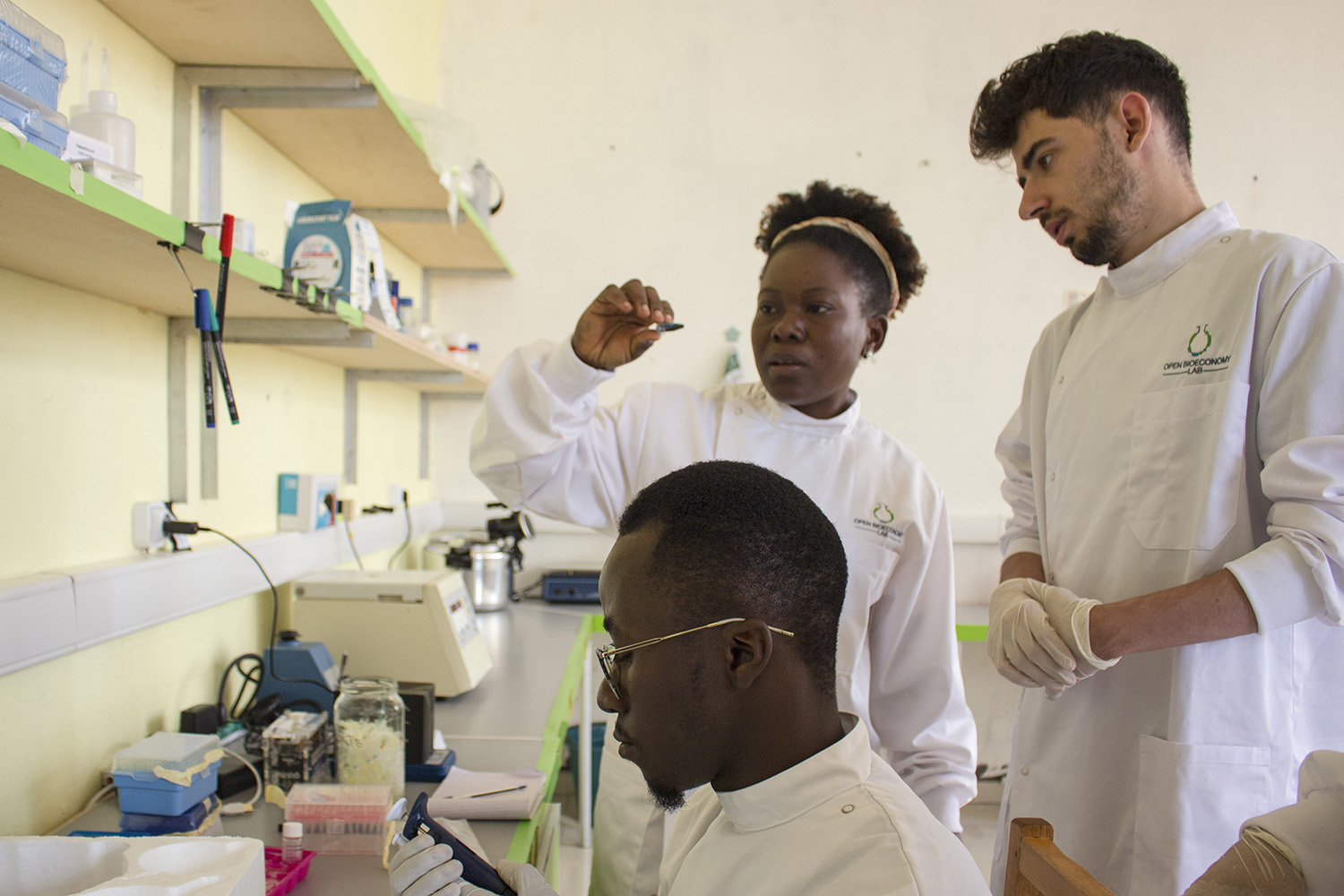
In 2016, Lenshina Agbor was working on her master’s thesis in Cameroon, analyzing the DNA of a parasite that causes malaria in pregnant women. She imported the enzymes needed for her research from Europe, but struggled with the cost of purchasing and shipping, months-long delays and the occasional batch of enzymes that became inactive during transport. As a result, she had to scale back her experiments.
Situations like Agbor’s are what inspired University of Cambridge researcher Jenny Molloy to contact Stanford bioengineer Drew Endy, who responded by creating an enzyme toolkit called Open Enzymes that makes biotechnology research more accessible to labs around the world.
“People everywhere now have the option of using these enzymes for free,” Endy said. “From students in middle school becoming literate in biotechnology to entrepreneurs creating a 21st-century DNA Valley, to citizens all over the world using biotechnology to solve local problems.”
With access to the Open Enzymes collection, Agbor and researchers like her can produce their own enzymes within days. “Since we have the Open Enzymes collection, upcoming researchers and diagnosticians would not have to go through some of the challenges I went through,” Agbor said.
Now, as she now works on her doctorate at Newcastle University Biosciences Institute in the U.K., Agbor said, “The gap in terms of research, research output, access to tools like enzymes and equipment couldn’t be wider.”
Narrowing that gap will allow scientists all over the world to carry out biotechnology research, said Keoni Gandall, a research assistant in Endy’s lab who worked on the enzyme set. “It’s a small step toward a world where everyone, regardless of race, gender, age or nation, can participate in biotechnology.”
Open-source science
One challenge for global labs lies in the fact that biotechnology supplies need continuous access to refrigeration – known as a cold chain – from when they leave a warehouse to when they arrive at a lab. “Enzymes for my teams in Ghana and Cameroon typically cost more than three times as much as in the U.S. and take weeks to months to arrive via an unreliable cold chain,” said Molloy. This concern is what led her to submit her request for a collection of enzymes that could solve all these problems to the BioBricks Foundation Free Genes Project, based out of Endy’s lab.
The Free Genes Project accepts requests for DNA sequences to be developed under an Open Materials Transfer Agreement (OpenMTA); once synthesized, the DNA is made available to the public, creating an open biotechnology commons. This service, founded by Endy, supports collaboration across international and institutional boundaries, promoting access in less privileged regions and institutions.
“Jenny and her team had done all the careful work to determine which of the most useful enzymes had come off patent,” said Endy, who is an associate professor of bioengineering. Free of claims on usage or materials, Endy could offer the enzymes for free to labs and researchers like Agbor.
Hardier enzymes
Endy said the enzymes in the collection are some of the basic workhorses of molecular biotechnology. “They are the molecular tools that let people read and edit DNA constructs,” he said. Until now, these critical enzymes had to be transported within a couple of days packed with dry ice.
“Those places that don’t have a cold chain can’t do the work. Period,” said Marc Fleischmann, administrative associate in Endy’s lab. Lack of access to these enzymes could prevent entire regions, especially developing nations, from participating in the biotech economy.
To overcome the need for refrigeration, the Endy lab genetically modified bacteria – which don’t need to be kept cold – to produce the enzymes. Once transported to a lab the bacteria can produce the enzymes locally for research purposes. As a result, distribution becomes simpler, less costly and more accessible to people worldwide.
“It’s super important to set up labs in developing countries to have the capability of local production,” Gandall said. “Otherwise it becomes prohibitively expensive to get the enzymes to the locations where they are needed.”
Global application
In addition to Cameroon, several community labs in the United States, Canada and Mexico, as well as Molloy’s partners in Ghana, have received the enzymes and are testing for DNA analysis. Molloy hopes the enzyme set will reach labs in Ethiopia, Chile, Argentina, India and Egypt in 2020.
In Chile, Tamara Matute Torres is eagerly awaiting the enzymes, which could help her sidestep bureaucratic complications when importing. Their arrival will allow her to carry out environmental monitoring of invasive aquatic species and potato viruses in her community.
“Access to tools and infrastructure is one concrete barrier that we know we can take steps to improve,” Molloy said. “The best use of the collection is probably the one we haven’t thought of yet.”
With the Open Enzymes collection, Agbor sees a brighter future of more than just research in Cameroon, but also the development of diagnostic tools for infectious diseases such as malaria, typhoid and hepatitis A. “This holds a lot of promise for me in terms of research quality and output,” Agbor said. She plans to return to Cameroon after her doctorate to take a research position in a local university.
Endy describes the increasing openness of biotechnology today like the construction of a language. The Open Enzymes collection is one step toward allowing synthetic biologists from anywhere in the world to join the conversation.
To read all stories about Stanford science, subscribe to the biweekly Stanford Science Digest.